Rather than being manufactured to 'look' or 'act' like antibodies, antibodies produced by phage display are the genuine article and support the requirement within Directive 2010/63/EU for a non-animal alternative to provide the same or higher level of information as the animal procedure...
Combinatorial diversification in nature
The recombination of gene segments to create the vast diversity of different antigen binding sites is an amazing phenomenon evolved by nature and unique to antibody genes: The relatively small number of genes dedicated to antibody production in the human genome would not otherwise be able to supply the enormous array of genetic material necessary to produce the vast diversity of different antibodies required to bind to all the potential antigens that theoretically could be encountered in the human body, some of which may only differ by as little as one amino acid. Each of the 3 immunoglobulin loci (IgG heavy, and IgG light [kappa and lambda chains) contains multiple copies of three different types of gene segments coding for the variable (binding) regions of the antibody molecule. These are known as Variable, Diverse, and Joining gene segments V(D)J. In a process called combinatorial diversification, or V(D)J recombination, DNA rearrangement causes one copy of each type of gene segment to be randomly recombined in any given lymphocyte. Combined with enzyme catalysed de novo DNA synthesis, an enormous antibody repertoire is generated with a huge molecular diversity at the antibody binding site. Roughly 3×1011 combinations are possible, although some are removed due to self-reactivity.
Creating diversity in a test tube
By exploiting this combinatorial mechanism, the same vast diversity of antibodies is achieved by phage display as well. For the construction of natural, naïve libraries, antibody encoding gene segments are simply copied from the DNA of human B lymphocytes (easily obtainable as cellular by-product of donor blood from human blood banks) and amplified by PCR using primers to the V regions. Alternatively, reliance on natural repertoires can be avoided completely by the use of synthetic libraries that are constructed by combinatorial mutagenesis with synthetic DNA technology. The natural or synthetic gene fragments are cloned into a phagemid vector and fused to minor coat protein III, which ensures antibodies will eventually be expressed on the surface of a phage particle, fused to the coat protein and thereby exposed for antigen recognition.
Packaging
Upon infection of the bacterial host E.coli, injection of the cloned phagemid into the cytoplasm, introduction of a helper phage to code for structural support and the subsequent provision of all genetic information required for assembly of new antibody expressing phage, an enormous diversity of antibody segments is generated. New phage particles are reproduced that display an antibody on their surface and one copy of the gene encoding it inside. Each phage therefore carries one specificity of binding activity, as well as exactly the right gene encoding for it. In this way, it is easy to have the entire human antibody repertoire, in a phage display library, ready for selection of the right binder. This process achieves a theoretical diversity of up to 1011 independent clones, equivalent to a life time’s supply of immunized animals.
Panning

Once the phage display library is constructed, selecting the right antibody to bind to the antigen from the enormous repertoire of possible binding site amino acid combinations might seem a daunting prospect, analogous to searching for the needle in the haystack. In fact, this selection process resembles that which takes place in nature and is similar, in practice, to that performed when isolating antibodies secreted by hybridomas. For animal derived monoclonal antibody production, a B lymphocyte population enriched by cells secreting antibodies to the target of interest, is isolated from the spleen of immunized animals and fused with myeloma cells to produce an immortal hybridoma cell. Each cell will only secrete antibodies specific for one epitope. Antibody candidates are selected by incubating secreted antibodies from each cell with the immobilised target of interest and good binders are subsequently expanded in vitro.
The phage display method is essentially the same except that animal immunization and fusion to myeloma cells is not a necessary step since the clonal selection completely takes place in bacteria and antibodies are expressed on the phage surface rather than secreted. Similarly to hybridoma cells, each phage in the library will express an antibody directed towards one epitope only. The selection is simply achieved by incubating the phage library suspension with the immobilised target of interest. The correct phage attaches to it via its surface displayed antibody and is retained while the others are washed away during consecutive rounds of ‘panning’. Careful design of the wash buffer eliminates binders that do not strongly associate with the target, ensuring that only strong binders are retained. Being a phage, this single attached antibody carrying its own gene in a backpack can be collected and used to reinfect a new E. coli cell, thus creating a monoclonal, antibody-of-choice producing cell line. There is one significant advantage with the in vitro based clonal selection using phage over the in vivo based clonal selection using hybridomas: Since the very moment of definition of antibody specificity is taking place in a test tube instead of the lymph node of an animal, the biochemical milieu can be controlled. This presents a significant advantage over animal based antibody generation, as the specificity can be defined and shaped from the start. For example, soluble competitors can be used to avoid cross-reactivities, or sequential pannings on related but slightly different antigens allows the functional identification of common epitopes.
Affinity maturation
During the course of a host’s immune response, following combinatorial diversification to produce germline antibodies, antibody affinity to the antigen continues to increase. The main principle of in vivo affinity maturation, namely somatic hypermutation followed by competitive clonal selection, is to produce mutations in the variable, antigen-binding coding sequences (complementarity-determining regions or CDRs) of the IgG genes, giving rise to a mutation rate up to a million times higher than in cell lines outside the lymphoid system. With repeated exposures to the same antigen, a host will produce antibodies of successively greater efficacy, generating antibodies with several log fold greater affinity.
Since the gene encoding the antibody is accessible, combinatorial based improvements are again, easily adopted by phage display, if required, to increase the affinity and specificity of the antibody selected during panning, completely in vitro. These include designed or PCR based random mutagenesis or variable chain shuffling as diversity creating mechanisms. They generate a vast repertoire of antibody variants, beyond that achievable by the host’s natural immune response, in consequence of accessibility to a much larger combinatorial space facilitating raising antibodies against basically any structure. Improvements in other antibody characteristics such as stability, pharmacokinetic parameters, effector parameters, tissue penetrating capacity of antibodies and their immunogenicity is possible by carefully adjusting the in vitro selection conditions.
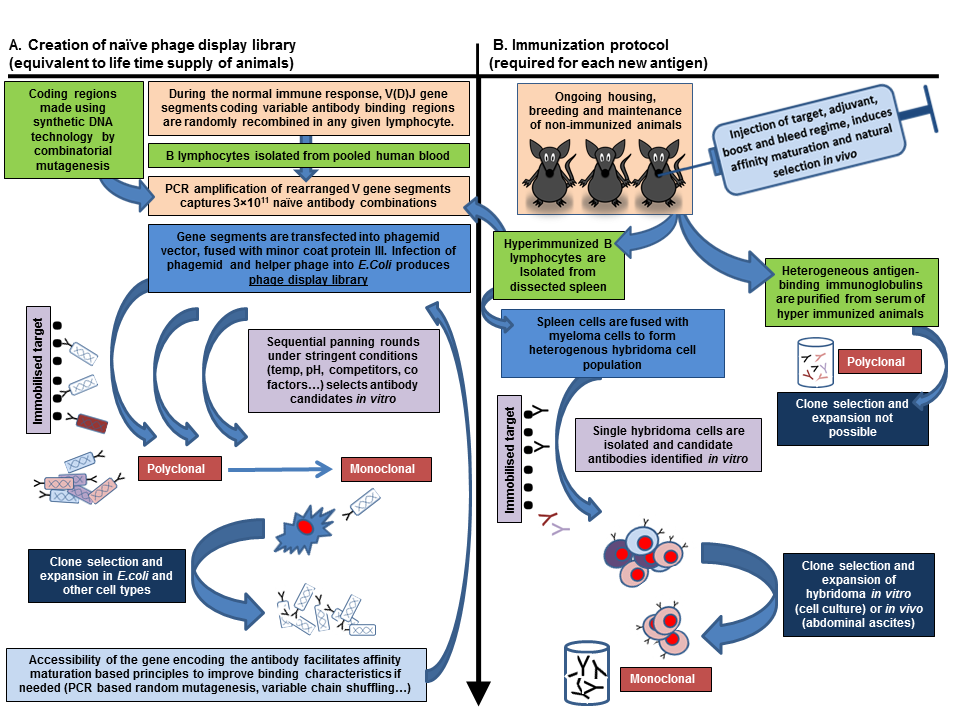
Schematic showing how the biological mechanisms employed by the in vivo adaptive immune system are adopted to produce phage display or animal derived antibodies (same colour boxes highlight areas of similarity)
Source: Gray et al (2016). Animal friendly affinity reagents - replacing the needless in the haystack.Trends in Biotechnology (in press)
Technical comparisons
Antibodies are frequently criticized for their poor quality, in particular their ability to recognize a defined target, their specificity for that target or ability to work in certain applications. A scientist purchasing an antibody from a commercial supplier is reliant upon them for quality and yet no standardised criteria exist, to which the supplier must adhere, to control its quality. Consequently, the burden of evidence upon which scientific reputation is staked, remains entirely with the scientist. The following table focusses on the scientific and practical differences between animal derived antibodies (monoclonal and polyclonal) and recombinant antibodies from naïve phage display libraries
Source: Gray et al (2016). Animal friendly affinity reagents - replacing the needless in the haystack.Trends in Biotechnology (in press)